DNA Sequencing Services:
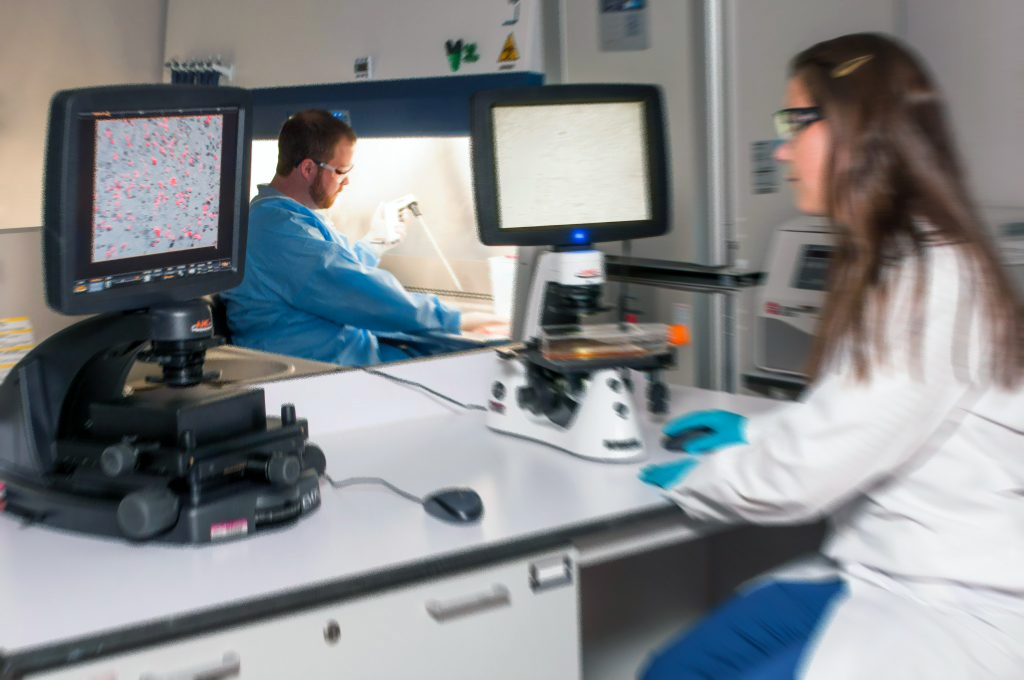
DNA sequencing is a means to determine the exact order of the building blocks of any genetic code. There are four subunits, also called nucleotide bases, that have been identified that make up all DNA. Those bases are: Adenine, Guanine, Cytosine, and Thymine. Knowledge pertaining to DNA sequences has become the next frontier for biological research. Being able to determine the sequence order can unlock vast amounts of information provided within a sample. Once this sequence information is gathered, it can be used to explore which stretches of DNA contain genes and which stretches carry regulatory instructions which can, for example, turn genes on and off. In addition, and most importantly, sequence data can highlight changes, routinely called mutations. Mutations in genes are mostly associated the precursors or predisposition to disease.
In recent years, as computer technology has rapidly evolved, so has the speed of DNA sequencing. Combined with major advances in molecular biology, modern DNA sequencing technology has been instrumental in the research and evolution of complete DNA sequences for numerous types and species of life, including the human genome and numerous animal, plant, and microbial species. What we do here at Fry Laboratories, L.L.C. is use this DNA sequence information to identify pathogens found in a clinical sample directly.



